Mag-Bind® TotalPure NGS
$0.00 – $4,272.40Price range: $0.00 through $4,272.40
Clean-up and size selection of DNA and RNA for NGS workflows using magnetic beads
- No protocol change against major competitor
- NGS: Double or single-sided size selection
- DNA cleanup: PCR cleanup
- RNA cleanup: cDNA or RNA purification
- Manual or adaptable to most open-ended liquid handlers
- Significant cost savings
- 96- or 384-well formats
The Mag-Bind® TotalPure NGS Kit allows for double or single-sided size selection. Purified PCR products can be used for myriad downstream applications such as microarrays, restriction enzyme digestion, and sequencing. Our Mag-Bind® magnetic bead-based technology provides an optimal alternative to centrifugation and vacuum-based extractions. This system is automatable on most open liquid handlers and magnetic processors.
Mag-Bind® TotalPure NGS offers an easy-to-use, reliable solution for the purification of DNA or RNA for Next Generation Sequencing workflows with high recovery rates. Mag-Bind TotalPure NGS is capable of selectively binding fragments depending on the reagent-to-sample ratio used, giving the user flexibility to perform left, right, or double-sided size selection. This product is designed for both manual or fully automated purification of DNA and RNA samples, as well as for the purification of PCR products. The system combines Omega Bio-tek’s proprietary chemistries with reversible nucleic acid binding properties of magnetic beads to selectively bind fragments larger than 100 bp and eliminate excess nucleotides, primers, and small, non-targeted products such as primer-dimers. Purified DNA and RNA are suitable for a variety of downstream applications such as NGS library preparation, microarrays, automated fluorescent sequencing, and restriction enzyme digestion.
For Research Use Only. Not for use in diagnostic procedures.
| FEATURES | SPECIFICATIONS |
|---|---|
| Downstream application | Cloning, NGS, In Vitro Transcription, Nucleic Acid Labeling, PCR, Real-Time Quantitative PCR (qPCR), Sequencing, Southern Blotting |
| Elution volume | 15 µL or above |
| Starting material | DNA or RNA: PCR products, gDNA, cDNA |
| Starting amount | Scalable |
| DNA recovered | >90% recovery for DNA >100 bp |
| Processing mode | Automated; manual |
| Throughput | 96-384 samples per run |
| DNA binding technology | Magnetic beads |
| Storage | 2°C – 8°C |
| Special note | Size selection by varying beads ratio |
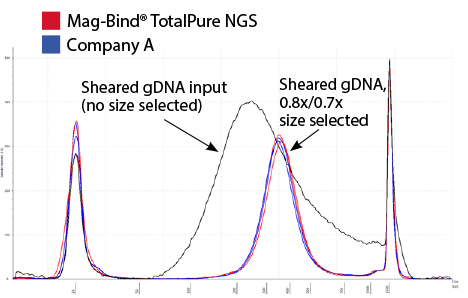
Double-Sided Selection
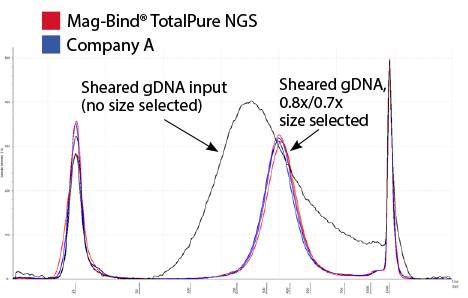
Figure 1. Electropherogram overlay of the double-sided size selection on sheared gDNA at 0.8x/0.7x ratio set using Omega Bio-tek’s Mag-Bind® TotalPure NGS and a comparable kit from Company A following manufacturer’s recommended protocols. The DNA was eluted in 25 µL and analyzed on Agilent’s TapeStation® 2200.
Recover Targeted DNA Fragments
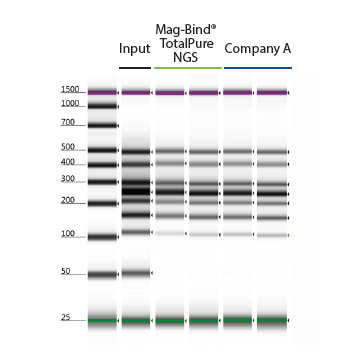
Figure 2. 10 µL of 50 bp ladder was purified with Omega Bio-tek’s Mag-Bind® TotalPure NGS and a comparable product from Company A following manufacturer’s recommended protocols. The DNA was eluted in 20 µL and analyzed on Agilent’s TapeStation® 2200.
Wide RNA Cleanup Concentration Range
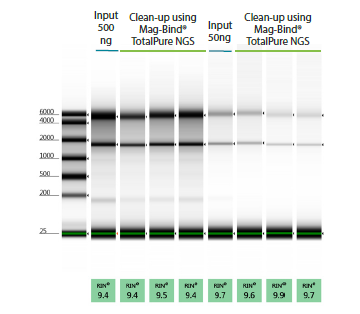
Figure 3. 10 µL of RNA at 50 ng/µL and 5 ng/µL was cleaned up with Omega Bio-tek’s Mag-Bind® TotalPure NGS following manufacturer’s recommended protocols. The RNA was eluted in 20 µL and analyzed on Agilent’s TapeStation® 2200. Recovery rates were 85-92% respectively.
Double-Sided Size Selection for NGS Library Prep
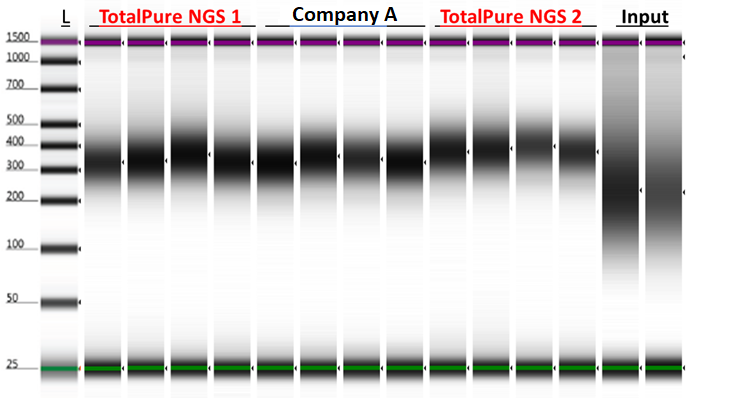
Figure 4. Next-generation sequencing libraries prepared from 350 ng sheared genomic DNA using Kapa Biosystem’s HyperPrep Kits (KK8504) and Omega Bio-tek’s Mag-Bind® TotalPure NGS and a comparable product from Company A on the Hamilton Microlab® STAR™. Mag-Bind® TotalPure NGS was used for 2 clean up step (0.8x and 1.0x) following Kapa Biosystems’ recommended protocol for clean up. DNA was analyzed on Agilent’s TapeStation® 2200 following library construction.
TapeStation Analysis of DNA Post-Cleanup at Different Bead-to-Sample Ratios

Figure 5. TapeStation analysis was performed on 25 μL of 20X diluted 50 bp ladder following cleanup with Mag-Bind TotalPure NGS beads on Tecan Fluent 780 workstation and an unprocessed 50 bp ladder as a control. The fragment sizes of DNA eluted after different cleanup ratios are as shown above.
Percentage of DNA Omitted at Different Bead:Sample Ratios

Figure 6. Average percentage of DNA removed (n=4) at various fragment sizes at different volume ratios of Mag-Bind TotalPure NGS beads to input sample volume. Cleanup was performed using the Tecan Fluent 780 workstation. Note: Negative percentages are reported as 0%.
Average Percentage of DNA Removed

Figure 7. Comparison of average percentage of DNA removed (n=4) at various bead:sample ratios following automated protocol on the Tecan Fluent 780 workstation to that of the data reported in [1]. Note: Negative percentages are reported as 0%.
- Evaluation of Omega Mag-Bind® TotalPure NGS Beads for DNA Size Selection. Genomics & Cell Characterization Core Facility, University of Oregon Technical Note – http://gc3fstorage.uoregon.edu/IMAGES/Evaluation_of_Omega_ Mag-Bind_TotalPure_NGS_Beads_MWeitzman_April2018.pdf.
FAQs
Can M1378 be used for “High Molecular Weight (HMW)” DNA?
Mag-Bind TotalPure NGS can be used to clean HMW gDNA. We have cleaned up fragments as large as 50-60 kb with good efficiency.
Can M1378 be used for DNA and RNA?
Yes. DNA and RNA can be cleaned up using TotalPure NGS.
What is the smallest size recovery using TotalPure NGS?
We have great success in recovering fragments down to 100 bp. However, we have had some success in recovering fragments down to 50 bp with a protocol modification. Please add 1.8 volumes of beads and 1 volume of isopropyl alcohol.
What is the difference between TotalPure NGS and RxnPure Plus?
TotalPure NGS has a much quicker magnetization time than the RxnPure particles, and TotalPure NGS particles are produced and quality controlled in an RNase-free environment allowing them to be used in RNA clean-up applications.
Product Information
| Name | Document | Languages | Link |
|---|---|---|---|
| Quick Guide | Product Manual | English | |
| Protocol Manual | Product Manual | English | |
| Mag-Bind TotalPure NGS | Product Literature | English | |
| Automated DNA Cleanup for PCR and NGS Workflows: Mag-Bind® TotalPure NGS on Tecan Fluent® 780 Workstation | Application Note | English | |
| In vitro Transcribed RNA Cleanup with Mag-Bind® TotalPure NGS | Application Note | English | |
| Evaluation of Omega Bio-tek Mag-Bind TotalPure NGS Beads | Application Note | English | |
| Removal of Genomic DNA Contamination from Purified cfDNA Using Mag-Bind® TotalPure NGS: A cfDNA Enrichment Methodology | Application Note | English |
Safety Data Sheets
| Components | Hazard Standards | Languages | Link |
|---|---|---|---|
| Mag-Bind TotalPure NGS | GHS | English | |
| Mag-Bind TotalPure NGS | GHS | Spanish | |
| Mag-Bind TotalPure NGS | REACH | English | |
| Mag-Bind TotalPure NGS | REACH | Danish | |
| Mag-Bind TotalPure NGS | REACH | Finnish | |
| Mag-Bind TotalPure NGS | REACH | French | |
| Mag-Bind TotalPure NGS | REACH | German | |
| Mag-Bind TotalPure NGS | REACH | Italian | |
| Mag-Bind TotalPure NGS | REACH | Norwegian | |
| Mag-Bind TotalPure NGS | REACH | Spanish | |
| Mag-Bind TotalPure NGS | REACH | Swedish | |
| Mag-Bind TotalPure NGS | WHMS | English | |
| Mag-Bind TotalPure NGS | WHMS | French | |
| Mag-Bind TotalPure NGS | REACH | Latvian | |
| Mag-Bind TotalPure NGS | REACH | Polish |