E.Z.N.A.® Soil DNA Kit
$0.00 – $839.90
- Reliable – Reproducible DNA purification from a variety of sample sources
- High quality – Ready-to-use DNA eliminating PCR inhibitors using proprietary inhibitor removal technology
- Yield – Efficient purification of DNA from even specialized samples
- Ease of use – Contains glass beads pre-filled in 2 mL vials
The E.Z.N.A.® Soil DNA Kit allows rapid and reliable isolation of high-quality genomic DNA from various soil samples. This kit can isolate microbial DNA from yeast, fungi, and grampositive or negative bacteria that inhabit a range of samples including clay, sandy, peaty, chalky, or loamy soil samples. This kit not only includes Disruptor Tubes prefilled with glass beads for efficient sample homogenization but also features a unique inhibitor removal reagent (cHTR Reagent) for effective removal of humic acid and other PCR inhibitors from eluted DNA. The extraction methodology combines the reversible nucleic acid-binding properties of HiBind® matrix with the speed and versatility of spin column technology for DNA purification. Up to 250 mg soil samples can be processed in 60 minutes or up to 1g soil samples in 2.5 hours. Purified DNA is suitable for PCR, restriction digestion, and next-generation sequencing.
For Research Use Only. Not for use in diagnostic procedures.
| FEATURES | SPECIFICATIONS |
|---|---|
| Starting Amount | Up to 1 g |
| Starting Material | Soil |
| Yield | Dependent upon sample |
| Elution Volume | 50-100 μL |
| Technology | HiBind® DNA Mini Column |
| Processing Mode | Manual |
| Throughput | 1-24 |
| ITEM | AVAILABLE SEPARATELY |
|---|---|
| HiBind® DNA Mini Columns | View Product |
| 2 mL Collection Tubes | View Product |
| Disruptor Tubes | View Product |
| SLX-Mlus Buffer | View Product |
| DS Buffer | Call for Pricing |
| P2 Buffer | View Product |
| cHTR Reagent | — |
| XP1 Buffer | — |
| HBC Buffer | View Product |
| DNA Wash Buffer | View Product |
| Elution Buffer | View Product |
Yield Comparison
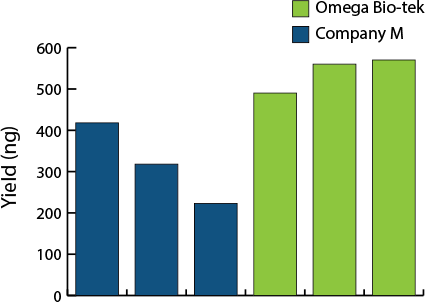

Figure 1. Comparison of DNA extraction method from soil samples. DNA yield determined with fluorescence-based dye quantification. 50 µL ZymoBIOMICS™ Microbial Community Standard was added to 200 mg soil samples and DNA was extracted using manufacturer’s recommended protocols. DNA was eluted in 100 µL for both manufacturers.
qPCR Comparison
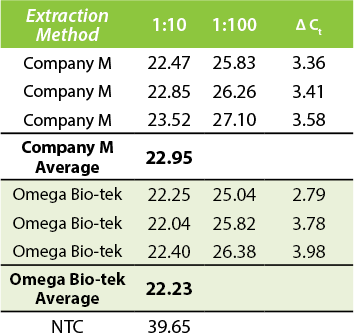

Figure 2. Comparison of Ct values. 20 µL SYBR Green qPCR reaction. 50 µL ZymoBIOMICS™ Microbial Community Standard was added to 200 mg soil samples and DNA was extracted using manufacturer’s recommended protocols. DNA was eluted in 100 µL for both manufacturers.
Bacteria Yield Comparison
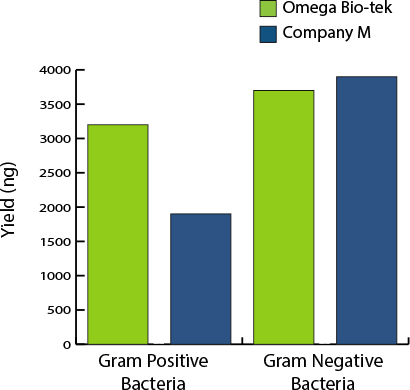

Figure 3. DNA yield by bacterial classes. DNA yield determined with fluorescence-based dye quantification. 0.5 mL cultured Gram-positive and Gram-negative bacteria were added to corresponding 200 mg soil samples and DNA was extracted using manufacturer’s recommended protocols. DNA was eluted in 100 µL for both manufacturers.
FAQs
I left my cHTR reagent out at room temperature for a few days. Is it ok to use?
We have stability data showing that the reagent is fine for three months at room temperature with no effect on quality. So it is ok to use. Please store at recommended temperature going forward.
I noticed precipitates in the cHTR buffer- is that normal? It is also difficult to pipette.
Precipitates are normal in cHTR. Please shake well before using and cut the pipette tip to help with aspiration.
Product Information
Safety Data Sheets
| Components | Hazard Standards | Languages | Link | hf:tax:dlp_document_language | hf:tax:dlp_document_hazard-standard |
|---|---|---|---|---|---|
| cHTR Reagent | GHS | English | english | ghs | |
| cHTR Reagent | GHS | Spanish | spanish | ghs | |
| cHTR Reagent | REACH | English | english | reach | |
| cHTR Reagent | REACH | Danish | danish | reach | |
| cHTR Reagent | REACH | Finnish | finnish | reach | |
| cHTR Reagent | REACH | French | french | reach | |
| cHTR Reagent | REACH | German | german | reach | |
| cHTR Reagent | REACH | Italian | italian | reach | |
| cHTR Reagent | REACH | Norwegian | norwegian | reach | |
| cHTR Reagent | REACH | Spanish | spanish | reach | |
| cHTR Reagent | REACH | Swedish | swedish | reach | |
| cHTR Reagent | WHMS | English | english | whms | |
| cHTR Reagent | WHMS | French | french | whms | |
| DNA Wash Buffer | GHS | English | english | ghs | |
| DNA Wash Buffer | GHS | Spanish | spanish | ghs | |
| DNA Wash Buffer | REACH | English | english | reach | |
| DNA Wash Buffer | REACH | Danish | danish | reach | |
| DNA Wash Buffer | REACH | Finnish | finnish | reach | |
| DNA Wash Buffer | REACH | French | french | reach | |
| DNA Wash Buffer | REACH | German | german | reach | |
| DNA Wash Buffer | REACH | Italian | italian | reach | |
| DNA Wash Buffer | REACH | Norwegian | norwegian | reach | |
| DNA Wash Buffer | REACH | Spanish | spanish | reach | |
| DNA Wash Buffer | REACH | Swedish | swedish | reach | |
| DNA Wash Buffer | WHMS | English | english | whms | |
| DNA Wash Buffer | WHMS | French | french | whms | |
| DS Buffer | GHS | English | english | ghs | |
| DS Buffer | GHS | Spanish | spanish | ghs | |
| DS Buffer | REACH | English | english | reach | |
| DS Buffer | REACH | Danish | danish | reach | |
| DS Buffer | REACH | Finnish | finnish | reach | |
| DS Buffer | REACH | French | french | reach | |
| DS Buffer | REACH | German | german | reach | |
| DS Buffer | REACH | Italian | italian | reach | |
| DS Buffer | REACH | Norwegian | norwegian | reach | |
| DS Buffer | REACH | Spanish | spanish | reach | |
| DS Buffer | REACH | Swedish | swedish | reach | |
| DS Buffer | WHMS | English | english | whms | |
| DS Buffer | WHMS | French | french | whms | |
| DS Buffer | REACH | Dutch | dutch | reach | |
| DS Buffer | REACH | Hungarian | hungarian | reach | |
| DS Buffer | REACH | Portuguese | portuguese | reach | |
| DS Buffer | REACH | Greek | greek | reach | |
| Elution Buffer | GHS | English | english | ghs | |
| Elution Buffer | GHS | Spanish | spanish | ghs | |
| Elution Buffer | REACH | English | english | reach | |
| Elution Buffer | REACH | Danish | danish | reach | |
| Elution Buffer | REACH | Finnish | finnish | reach | |
| Elution Buffer | REACH | French | french | reach | |
| Elution Buffer | REACH | German | german | reach | |
| Elution Buffer | REACH | Italian | italian | reach | |
| Elution Buffer | REACH | Norwegian | norwegian | reach | |
| Elution Buffer | REACH | Spanish | spanish | reach | |
| Elution Buffer | REACH | Swedish | swedish | reach | |
| Elution Buffer | WHMS | English | english | whms | |
| Elution Buffer | WHMS | French | french | whms | |
| Elution Buffer | REACH | Dutch | dutch | reach | |
| Elution Buffer | REACH | Hungarian | hungarian | reach | |
| Elution Buffer | REACH | Portuguese | portuguese | reach | |
| Elution Buffer | REACH | Greek | greek | reach | |
| HBC Buffer | GHS | English | english | ghs | |
| HBC Buffer | GHS | Spanish | spanish | ghs | |
| HBC Buffer | REACH | English | english | reach | |
| HBC Buffer | REACH | Danish | danish | reach | |
| HBC Buffer | REACH | Finnish | finnish | reach | |
| HBC Buffer | REACH | French | french | reach | |
| HBC Buffer | REACH | German | german | reach | |
| HBC Buffer | REACH | Italian | italian | reach | |
| HBC Buffer | REACH | Norwegian | norwegian | reach | |
| HBC Buffer | REACH | Spanish | spanish | reach | |
| HBC Buffer | REACH | Swedish | swedish | reach | |
| HBC Buffer | WHMS | English | english | whms | |
| HBC Buffer | WHMS | French | french | whms | |
| P2 Buffer | GHS | English | english | ghs | |
| P2 Buffer | GHS | Spanish | spanish | ghs | |
| P2 Buffer | REACH | English | english | reach | |
| P2 Buffer | REACH | Danish | danish | reach | |
| P2 Buffer | REACH | Finnish | finnish | reach | |
| P2 Buffer | REACH | French | french | reach | |
| P2 Buffer | REACH | German | german | reach | |
| P2 Buffer | REACH | Italian | italian | reach | |
| P2 Buffer | REACH | Norwegian | norwegian | reach | |
| P2 Buffer | REACH | Spanish | spanish | reach | |
| P2 Buffer | REACH | Swedish | swedish | reach | |
| P2 Buffer | WHMS | English | english | whms | |
| P2 Buffer | WHMS | French | french | whms | |
| SLX M1us Buffer | GHS | English | english | ghs | |
| SLX M1us Buffer | GHS | Spanish | spanish | ghs | |
| SLX M1us Buffer | REACH | English | english | reach | |
| SLX M1us Buffer | REACH | Danish | danish | reach | |
| SLX M1us Buffer | REACH | Finnish | finnish | reach | |
| SLX M1us Buffer | REACH | French | french | reach | |
| SLX M1us Buffer | REACH | German | german | reach | |
| SLX M1us Buffer | REACH | Italian | italian | reach | |
| SLX M1us Buffer | REACH | Norwegian | norwegian | reach | |
| SLX M1us Buffer | REACH | Spanish | spanish | reach | |
| SLX M1us Buffer | REACH | Swedish | swedish | reach | |
| SLX M1us Buffer | WHMS | English | english | whms | |
| SLX M1us Buffer | WHMS | French | french | whms | |
| XP1 Buffer | GHS | English | english | ghs | |
| XP1 Buffer | GHS | Spanish | spanish | ghs | |
| XP1 Buffer | REACH | English | english | reach | |
| XP1 Buffer | REACH | Danish | danish | reach | |
| XP1 Buffer | REACH | Finnish | finnish | reach | |
| XP1 Buffer | REACH | French | french | reach | |
| XP1 Buffer | REACH | German | german | reach | |
| XP1 Buffer | REACH | Italian | italian | reach | |
| XP1 Buffer | REACH | Norwegian | norwegian | reach | |
| XP1 Buffer | REACH | Spanish | spanish | reach | |
| XP1 Buffer | REACH | Swedish | swedish | reach | |
| XP1 Buffer | WHMS | English | english | whms | |
| XP1 Buffer | WHMS | French | french | whms |

